Keeping drugs on the job
Computer simulations developed at the University of Michigan reveal how well drug additives stop the active ingredients from crystallizing in the digestive tract.
Computer simulations developed at the University of Michigan reveal how well drug additives stop the active ingredients from crystallizing in the digestive tract.
Computer simulations developed at the University of Michigan reveal how well drug additives stop the active ingredients from crystallizing in the digestive tract. They tested these simulations on the anti-seizure drug phenytoin.
“Most drugs are hydrophobic, so the mix of water and drugs is not stable,” said Taraknath Mandal, a research fellow in chemical engineering. “They spontaneously form a crystal.”
In the water-based environment of the stomach and small intestine, the active ingredients stick together, and then they don’t make it into the bloodstream. Pharma companies add polymers to drugs to get in the way of crystal growth, but the range of possible designs for these polymers is huge.
The computer simulations enable researchers to try out different polymers, looking for the best performance.
“These tools have never been used before on this problem,” said Ron Larson, the A. H. White Distinguished University Professor of Chemical Engineering and the George Granger Brown Professor of Chemical Engineering, who led the work. He and his team worked closely with researchers at Dow Chemical Company in Midland, Michigan, including WW (Trey) Porter, the team lead on the Dow side.
They tested the simulation with several drugs, focusing on how phenytoin interacted with a cellulose-based polymer – the stuff that makes up the cell walls in plants. They compared how different side groups on the cellulose chain affected how well the polymer kept phenytoin from crystallizing in the simulated gut environment.
The team tested two side-groups head to head – succinyl and acetyl. Both are needed to make a polymer effective in both the acidic environment of the stomach and the more neutral environment of the small intestine.
In the stomach, the succinyl groups have the full complement of hydrogen atoms, so they are not charged. They help the polymer interact with a hydrophobic drug like phenytoin. A charged polymer will be attracted to the water rather than the drug.
But once the polymers hit the small intestine, some hydrogen atoms strike out on their own, leaving the succinyl with a negative charge. Here, the acetyl groups take over helping to keep the polymer groups together even as the neutral succinyl groups expand. Larson and his group developed computer models to find the best balance.
Of course, patients don’t gain anything if the drug additives keep the drug from crystallizing but instead traps it in a web of polymers. Or, if the drugs are released too quickly, they can crystallize in the small intestine. Wenjun Huang, a doctoral student in chemical engineering, created a second simulation to test how well drugs leave the polymer webs.
“Our simulation results are the first ones to directly show the role of each individual functional group in the polymer,” said Huang. “We can see the polymer-drug aggregates form, and we can see how drugs are released from the aggregates.”
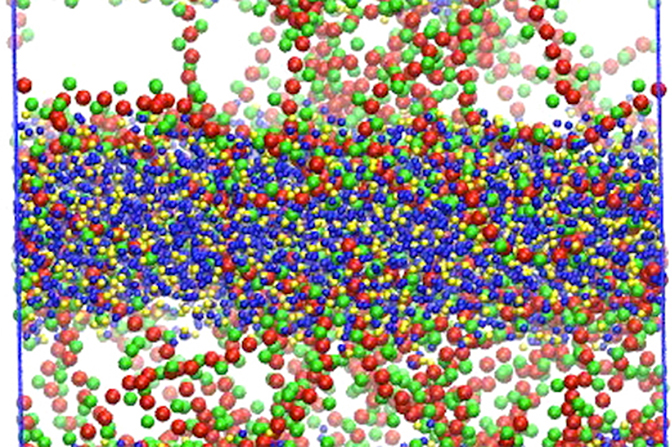
Again, a balance between the succinyl and acetyl groups is needed to achieve the right pace, and the balance is different for different drugs.
“We are beginning to learn how to determine computationally which modifications to the polymer will be most effective at interacting with a given drug and thereby help in the design of better polymers to enhance drug delivery,” said Larson.
These studies were funded primarily by the Dow Chemical Company. The team also relied on Advanced Research Computing at U-M, funded by the National Science Foundation.
The paper on the model that investigates how well different polymers work to prevent drug crystallization is titled “A framework for multi-scale simulation of crystal growth in the presence of polymers,” and it was published in Soft Matter.
The paper on the computer model that explores whether drugs are released from the web of polymers is titled “Computational Modeling of Hydroxypropyl-Methylcellulose Acetate Succinate (HPMCAS) and Phenytoin Interactions: A Systematic Coarse-Graining Approach,” and it was published in the journal Molecular Pharmaceutics.
Larson is also a professor of macromolecular science and engineering, biomedical engineering, and mechanical engineering, and is a member of the Biointerfaces program.